可拉伸细胞基底硬度控制培养皿(CellSoft 100mm Round Dishes)
Cells sense soft! CellSoft offers softer substrates to match the material properties of tissue niches to better meet the needs of biological laboratories wanting to grow their cells on native stiffness。
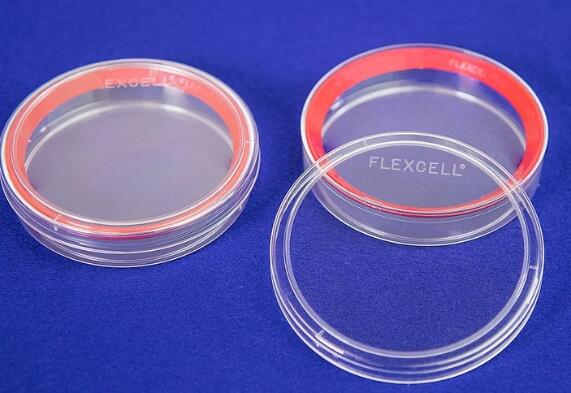
直径100mm培养皿,总生长表面积为57cm2
●BioFlex® CellSoft标准6孔板
●腔室载玻片CellSoft
CellSoft培养板有很多不同的种类,如不同的硬度,不同的孔板,用于显微观察的腔室载玻片(圆形多孔板),共价包被CollagenI或其他蛋白,可对细胞进行静态或动态牵拉应力刺激。更重要的一点,新型的CellSoft培养板可以反复酶消化和再接种细胞,蛋白包被的表面可以重复使用多达三次。
niche。●弹性模量范围1-80kPa
●BioFlex® CellSoft标准6孔板
●腔室载玻片CellSoft
Amino,
Elastin,
and Laminin (YIGSR)
and untreated (未处理)
纳米图案化牵张、压缩培养表面提供细胞微环境,模仿天然细胞外基质的对齐结构,促进细胞结构和功能发展。
PUBLICATIONS
Confinement and Low Adhesion Induce Fast Amoeboid Migration of Slow Mesenchymal Cells
Y.-J. Liu, M. Piel, Cell, et al., 2015 160(4), 659-672Actin flows induce a universal coupling between cell speed and cell persistence
P. Maiuri, R. Voituriez, et al., Cell, 2015 161(2), 374–386Geometric friction directs cell migration
M. Le Berre, M. Piel, et al., Physical Review Letter 2013 111, 198101Mitotic rounding alters cell geometry to ensure efficient spindle assembly
O. M. Lancaster, B. Baum, et al., Developmental Cell, 2013 25(3), 270-283Fine Control of Nuclear Confinement Identifies a Threshold Deformation leading to Lamina Rupture and Induction of Specific Genes
M. Le Berre, J. Aubertin, M. Piel, Integrative Biology, 2012 4 (11), 1406-1414Exploring the Function of Cell Shape and Size during Mitosis
C. Cadart, H. K. Matthews, et al., Developmental Cell, 2014 29(2), 159-169Methods for Two-Dimensional Cell Confinement
M. Le Berre, M. Piel, et al., 2014, Micropatterning in Cell Biology Part C, Methods in cell biology, 121, 213-29
纳米图案化牵张、压缩培养表面提供细胞微环境,模仿天然细胞外基质的对齐结构,促进细胞结构和功能发展。
References
[1] D. Huh, G.A. Hamilton, and D. E. Ingber, “From 3D cell culture to organs-on-chips,” TrendsCell Biol., vol. 21, no. 12, pp. 745–754, 2011.
[2] M. Ravi, V.Paramesh, S. R. Kaviya, E. Anuradha, and F. D. Paul Solomon, “3D cell culturesystems: Advantages and applications,” J. Cell. Physiol., vol. 230,no. 1, pp. 16–26, 2015.
[3] J. W.Haycock, 3D cell culture: a review of current approaches andtechniques., vol. 695. 2011.
[4] F.Pampaloni, E. G. Reynaud, and E. H. K. Stelzer, “The third dimension bridgesthe gap between cell culture and live tissue.,” Nat. Rev. Mol. CellBiol., vol. 8, no. 10, pp. 839–845, 2007.
[5] J. Lee, M.J. Cuddihy, and N. A. Kotov, “Three-dimensional cell culture matrices: state ofthe art.,” Tissue Eng Part B Rev, vol. 14, no. 1, pp. 61–86, 2008.
[6] M.Vinci et al., “Advances in establishment and analysis ofthree-dimensional tumor spheroid-based functional assays for target validationand drug evaluation,” BMC Biol., vol. 10, no. 1, p. 29, 2012.
[7] B. A.Justice, N. A. Badr, and R. A. Felder, “3D cell culture opens new dimensions incell-based assays,” Drug Discov. Today, vol. 14, no. 1–2, pp.102–107, 2009.
[8] I.Meyvantsson and D. J. Beebe, “Cell culture models in microfluidicsystems.,” Annu. Rev. Anal. Chem., vol. 1, pp. 423–449, 2008.
[9] E. W. K.Young and D. J. Beebe, “Fundamentals of microfluidic cell culture in controlledmicroenvironments,” Chem Soc Rev, vol. 39, no. 3, pp. 1036–1048,2010.
[10] D. J.Beebe, G. a Mensing, and G. M. Walker, “Physics and applications ofmicrofluidics in biology.,” Annu. Rev. Biomed. Eng., vol. 4, pp.261–286, 2002.
[11] J. El-Ali,P. K. Sorger, and K. F. Jensen, “Cells on chips.,” Nature, vol.442, no. 7101, pp. 403–411, 2006.
[12] J.Guck et al., “Optical deformability as an inherent cell marker fortesting malignant transformation and metastatic competence,” Biophys J,vol. 88, no. 5, pp. 3689–3698, 2005.
[13] S.Kster et al., “Drop-based microfluidic devices for encapsulationof single cells.,” Lab Chip, vol. 8, no. 7, pp. 1110–1115, 2008.
[14] H.Andersson and A. Van den Berg, “Microfluidic devices for cellomics: Areview,” Sensors Actuators, B Chem., vol. 92, no. 3, pp. 315–325,2003.
[15] M. W.Tibbitt and K. S. Anseth, “Hydrogels as extracellular matrix mimics for 3D cellculture,” Biotechnol. Bioeng., vol. 103, no. 4, pp. 655–663, 2009.
[16] J. P.Vacanti and R. Langer, “Tissue engineering: the design and fabrication ofliving replacement devices for surgical reconstruction andtransplantation.,” Lancet, vol. 354, p. SI32-I34, 1999.
[17] G. S. D.Hetal Patel, Minal Bonde, “Biodegradable polymer scaffolds for tissueengineering,” Trends Biomater. Artif. Organs, vol. 25, no. 1, pp.20–29, 2011.
[18] L. G.Griffith and M. A. Swartz, “Capturing complex 3D tissue physiology invitro.,” Nat. Rev. Mol. cell Biol., vol. 7, no. 3, pp. 211–24,2006.
[19] D. J.Tobin, “Scaffolds for Tissue Engineering and 3D Cell Culture,” MethodsMol. Biol., vol. 695, no. 2, pp. 213–227, 2011.
[20] J.Naranda et al., “Polyester type polyHIPE scaffolds with an interconnectedporous structure for cartilage regeneration,” Sci. Rep., vol. 6,no. February, p. 28695, 2016.
[21] B.Dhandayuthapani, Y. Yoshida, T. Maekawa, and D. S. Kumar, “Polymeric scaffoldsin tissue engineering application: A review,” Int. J. Polym. Sci.,vol. 2011, no. ii, 2011.
[22] F. J.O’Brien, “Biomaterials & scaffolds for tissue engineering,” Mater.Today, vol. 14, no. 3, pp. 88–95, 2011.
[23] A. L.Paguirigan and D. J. Beebe, “Microfluidics meet cell biology: Bridging the gap byvalidation and application of microscale techniques for cell biologicalassays,” BioEssays, vol. 30, no. 9, pp. 811–821, Sep. 2008.
[24] F.-Q. Nie,M. Yamada, J. Kobayashi, M. Yamato, A. Kikuchi, and T. Okano, “On-chip cellmigration assay using microfluidic channels.,” Biomaterials, vol.28, no. 27, pp. 4017–4022, 2007.
[25] A. Valster,N. L. Tran, M. Nakada, M. E. Berens, A. Y. Chan, and M. Symons, “Cell migrationand invasion assays,” Methods, vol. 37, no. 2, pp. 208–215, 2005.
[26] C. R.Justus, N. Leffler, M. Ruiz-Echevarria, and L. V Yang, “In vitro cell migrationand invasion assays.,” J. Vis. Exp., vol. 752, no. 88, p. e51046,2014.
[27] N.Kramer et al., “In vitro cell migration and invasionassays.,” Mutat Res, vol. 752, no. 1, pp. 10–24, 2013.
[28] J. W. Hong,V. Studer, G. Hang, W. F. Anderson, and S. R. Quake, “A nanoliter-scale nucleicacid processor with parallel architecture.,” Nat. Biotechnol., vol.22, no. 4, pp. 435–439, 2004.
[29] J. Q.Boedicker, L. Li, T. R. Kline, and R. F. Ismagilov, “Detecting bacteria anddetermining their susceptibility to antibiotics by stochastic confinement innanoliter droplets using plug-based microfluidics.,” Lab Chip, vol.8, no. 8, pp. 1265–1272, 2008.
[30] G.Velve-Casquillas, M. Le Berre, M. Piel, and P. T. Tran, “Microfluidic tools forcell biological research,” Nano Today, vol. 5, no. 1. pp. 28–47,2010.
[31] C. R.Terenna et al., “Physical Mechanisms Redirecting Cell Polarity andCell Shape in Fission Yeast,” Curr. Biol., vol. 18, no. 22, pp.1748–1753, . 2008.
[32] G.Faure-andré, “Regulation of Dendritic Cell Migration by CD74, the MHC ClassII–Associated Invariant Chain,” Science (80-. )., vol. 1705, no.December, 2008.
[33] S. M.McFaul, B. K. Lin, and H. Ma, “Cell separation based on size and deformabilityusing microfluidic funnel ratchets,” Lab Chip, vol. 12, no. 13, pp.2369–2376, 2012.
[34] S. C. Hur,N. K. Henderson-MacLennan, E. R. B. McCabe, and D. Di Carlo,“Deformability-based cell classification and enrichment using inertialmicrofluidics.,” Lab Chip, vol. 11, no. 5, pp. 912–920, 2011.
[35] H. W. Hou,Q. S. Li, G. Y. H. Lee, A. P. Kumar, C. N. Ong, and C. T. Lim, “Deformabilitystudy of breast cancer cells using microfluidics,” Biomed. Microdevices,vol. 11, no. 3, pp. 557–564, 2009.
我公司专注生物力学和生物打印等生物医学工程科研服务-10年经验支持,
点击查更多科研工具-应用尽有




















